Data-driven decomposition of neurophysiological event waveform populations.
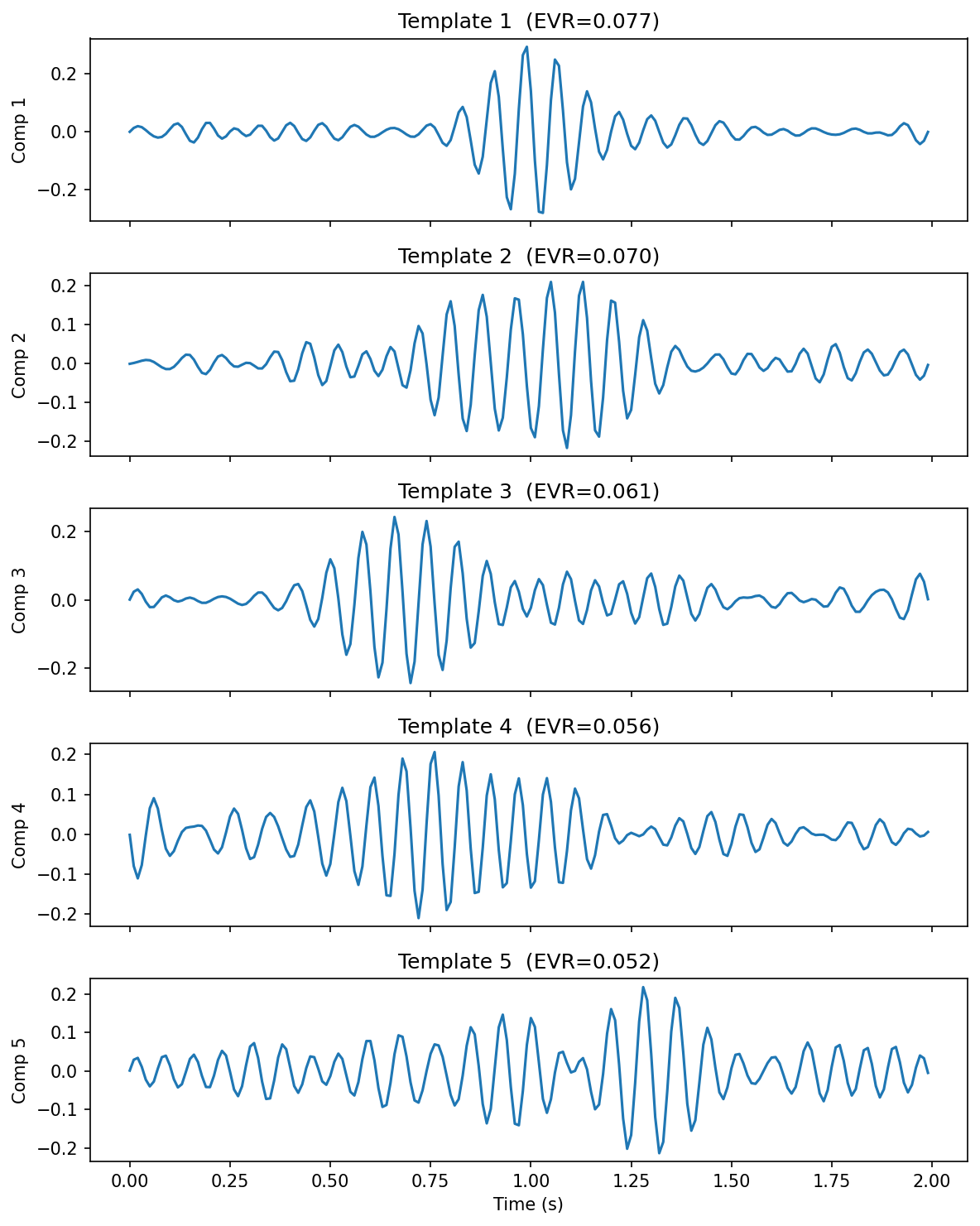
Five principal waveform templates extracted from 821 sleep spindles via SVD.
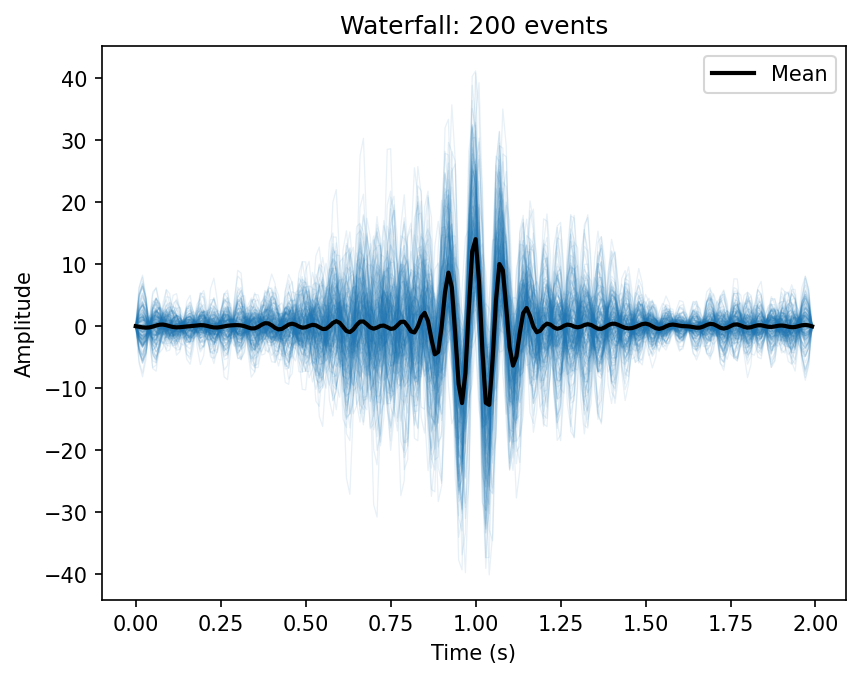
200 sigma-filtered spindle waveforms overlaid, with the population mean in black.
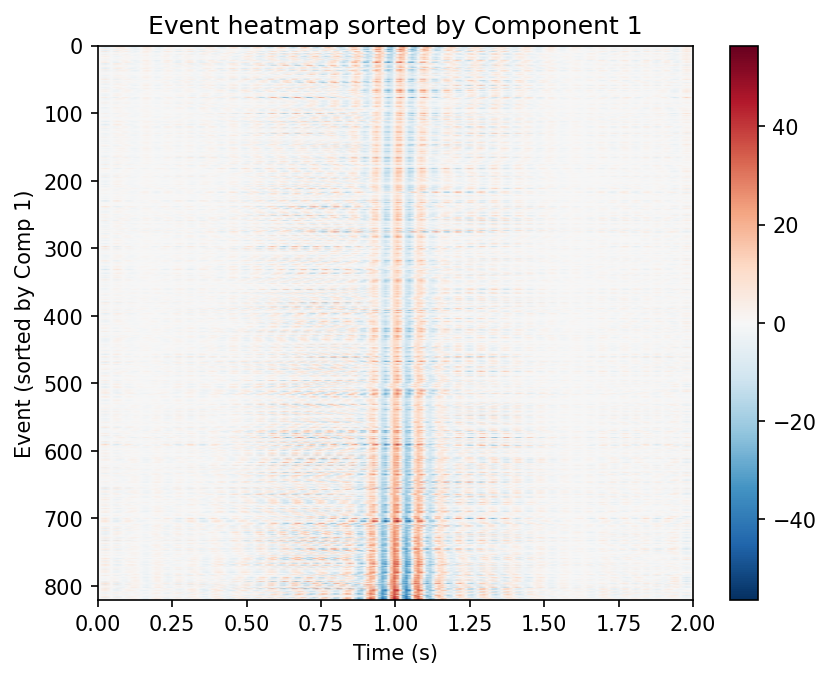
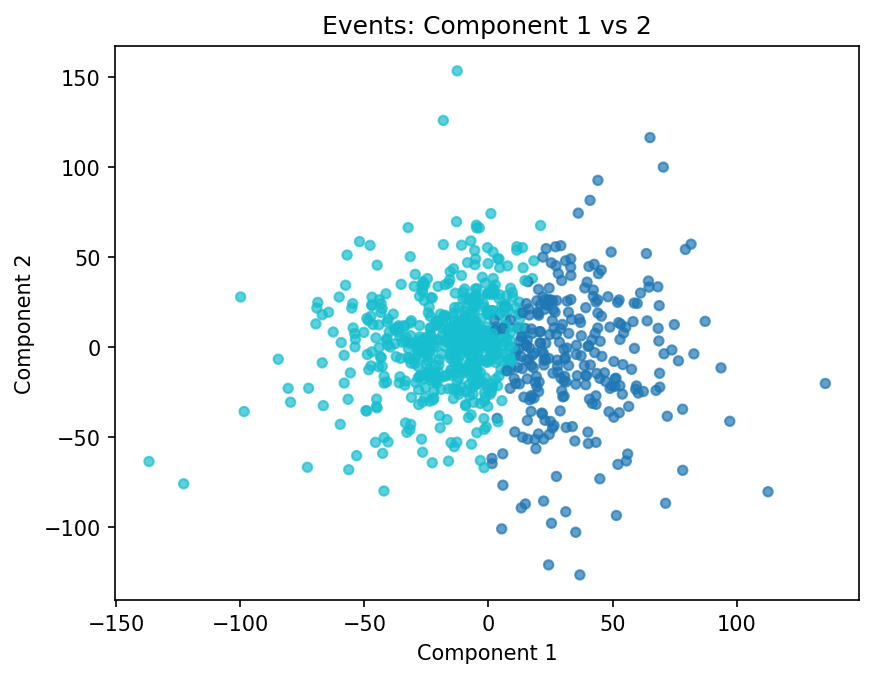
Left: all events sorted by Component 1 loading. Right: k-means clustering in the first two component scores.
pip install subwaveOptional extras:
pip install "subwave[mne]" # MNE-Python and Luna I/O
pip install "subwave[yasa]" # YASA spindle detection I/O (pulls in mne)
pip install "subwave[scattering]" # Wavelet scattering decomposition (kymatio)
pip install "subwave[tensor]" # Multi-channel tensor decomposition (tensorly)
pip install "subwave[all]" # All optional backendsimport numpy as np
import subwave as sw
rng = np.random.default_rng(0)
t = np.linspace(0, 1, 256)
X = np.stack([np.sin(2 * np.pi * 13 * t) + rng.normal(0, 0.1, 256) for _ in range(100)])
result = sw.decompose(X, method="svd", n_components=5)
result.plot_templates(n=3)Each template is a basis waveform shape; each loading is how strongly a given event expresses that template. Together they reconstruct the original population.
Or let subwave choose the number of components:
result = sw.decompose(X, n_components="auto")import numpy as np
import subwave as sw
# Stack detected QRS complexes into an event matrix
X_beats = np.load("heartbeats.npy") # (n_beats, n_samples)
result = sw.decompose(X_beats, method="svd", n_components=3)
# Outlier detection flags ectopic beats
scores = result.outlier_scores()
ectopic = np.where(scores > np.percentile(scores, 99))[0]sw.from_array(X, sfreq=256) # plain numpy
sw.from_npz("spindles.npz") # Lunascope format
sw.from_mne(epochs) # MNE Epochs
sw.from_yasa(spindles_df, raw_signal, sfreq=256) # YASA output
sw.from_luna("spindles.txt", "recording.edf", sfreq=256) # Luna output + EDF
sw.from_edf_batch(["s1.edf", "s2.edf"], channel="C3") # detect + pool across filesfrom_edf_batch runs a detector (default YASA) on every EDF, extracts windows
around each peak, and returns a single EventMatrix with a .meta DataFrame
recording subject, file, event_index, and peak_sec for each pooled
event.
- SVD / PCA (
method='svd') — default, optimal low-rank approximation. Uses randomized SVD for >5000 events. - NMF (
method='nmf') — parts-based, requires non-negative input. - Dictionary learning (
method='dictlearn') — sparse atoms. - Fourier-then-SVD (
method='fourier_svd') — shift-invariant: SVD on per-event magnitude rFFT spectra. Robust to small temporal jitter that corrupts plain SVD. Templates live in frequency space (config['domain'] = 'frequency'). - Scattering-then-SVD (
method='scattering_svd') — locally translation-invariant and stable to deformation via 1-D wavelet scattering (kymatio). Templates live in scattering-coefficient space. Requiressubwave[scattering].
For 3-D event tensors (n_events, n_samples, n_channels), tensor_decompose
factors temporal and spatial structure separately instead of flattening:
result = sw.tensor_decompose(X, method="cp", rank=3) # PARAFAC / CP
result = sw.tensor_decompose(X, method="tucker", rank=3) # Tucker (with core)
result.event_factors # (n_events, rank)
result.temporal_factors # (n_samples, rank)
result.spatial_factors # (n_channels, rank)
result.reconstruction_error # ||X - X_hat|| / ||X||
result.component_waveform(0, channel=1) # template scaled by channel weightRequires subwave[tensor].
result.templates # (n_components, n_samples) basis waveforms
result.loadings # (n_events, n_components) per-event scores
result.explained_variance_ratio # variance captured per component
result.singular_values # singular values
result.factor_tables["instance"] # DataFrame: instance_id, score_1…k, recon_error
result.reconstruct(n_components=3) # rank-k reconstruction
result.project(new_X) # project new events onto learned subspace
result.outlier_scores() # per-event reconstruction errork = sw.parallel_analysis(X) # Horn's parallel analysis
k = sw.elbow(result.singular_values) # Kneedle elbow detection
k = sw.kaiser(result) # Kaiser rulefreqs, powers = result.template_spectrum(sfreq=256)
peak_hz = result.template_peak_freq(sfreq=256) # e.g. [13.2, 11.1] Hz
bw_hz = result.template_bandwidth(sfreq=256)cr = result.cluster(method="kmeans", n_clusters=2)
cr["labels"] # cluster assignments
result.cluster_templates(n_clusters=2) # mean waveform per clusterperm = sw.permutation_test(X, groups, n_components=3, n_perm=500)
perm.p_value # do two groups span different subspaces?
df = sw.comparison.loading_test(result, groups, n_perm=1000)
# Columns: component, observed_diff, p_value, cohens_d, p_corrected
# cohens_d: standardized effect size (pooled SD)
# p_corrected: Benjamini-Hochberg FDR-adjusted p-values across componentsConvenience routines for sleep-spindle waveforms (sigma-band filtering, envelope-based alignment, canonical templates).
from subwave.spindles import (
sigma_filter, # 9–16 Hz Butterworth bandpass (sosfiltfilt)
align_by_envelope_peak, # circularly shift so each event's sigma-envelope peak is centered
CANONICAL_FAST, # ~13.5 Hz Gaussian-windowed sinusoid (256 Hz, 1 s)
CANONICAL_SLOW, # ~11 Hz Gaussian-windowed sinusoid (256 Hz, 1 s)
)
X_filt = sigma_filter(X, sfreq=256)
X_aligned = align_by_envelope_peak(X, sfreq=256)Tools for checking decompositions against ground truth and for estimating component reliability.
from subwave.validation import (
synthetic_population, # generate events from known templates with noise / jitter / amplitude variability
recovery_score, # cosine similarity between true and recovered templates (Hungarian-matched)
cluster_recovery_ari, # Adjusted Rand Index between true and recovered cluster labels
bootstrap_stability, # mean cosine similarity of bootstrap-resampled templates to full-data templates
split_half, # template & loading reproducibility on odd/even split
)
# Ground-truth recovery on synthetic data
truth = synthetic_population(templates, n_events=200, noise_std=0.1, random_state=0)
result = sw.decompose(truth["X"], method="svd", n_components=truth["templates"].shape[0])
score = recovery_score(truth["templates"], result.templates)
score["mean_score"] # cosine similarity (1.0 = perfect)
# Reliability of a real decomposition
boot = bootstrap_stability(X, n_components=3, n_boot=100, random_state=0)
boot["stability_scores"] # per-component mean similarity across bootstraps
sh = split_half(X, n_components=3, random_state=0)
sh["template_similarity"] # per-component template reproducibility
sh["loading_correlation"] # correlation of projected vs native loadingsresult.save("result.npz")
result = sw.load_result("result.npz")
df = result.to_dataframe() # flat DataFrame for R/Stataresult.plot_spectrum() # singular value scree plot
result.plot_templates(n=5) # basis waveforms
result.plot_template_spectra(sfreq=256) # power spectrum of each template
result.plot_scatter(x=0, y=1) # component 0 vs 1 (supports color= for clusters)
result.plot_heatmap(comp=0) # events × samples sorted by loading
result.plot_waterfall(n=100) # overlaid waveforms with bold mean
result.plot_mean_pm(comp=0) # mean ± component
result.plot_sorted_grid(comp=0) # events sorted by score
result.plot_residual_hist() # reconstruction error distribution
result.plot_cumulative_variance() # cumulative EVR curve
result.plot_reconstruction(event_idx=0) # original vs reconstruction
result.plot_loadings_by_group(groups) # box/violin by group
result.plot_loadings_over_time(times) # loading drift across timeSee docstrings via help(sw.decompose) for full options, including cluster_sweep, loading_test, subspace_angles, scatter_colored_by, loadings_correlated_with, and more.
If you use subwave in published work, see CITATION.cff in the repository.
MIT