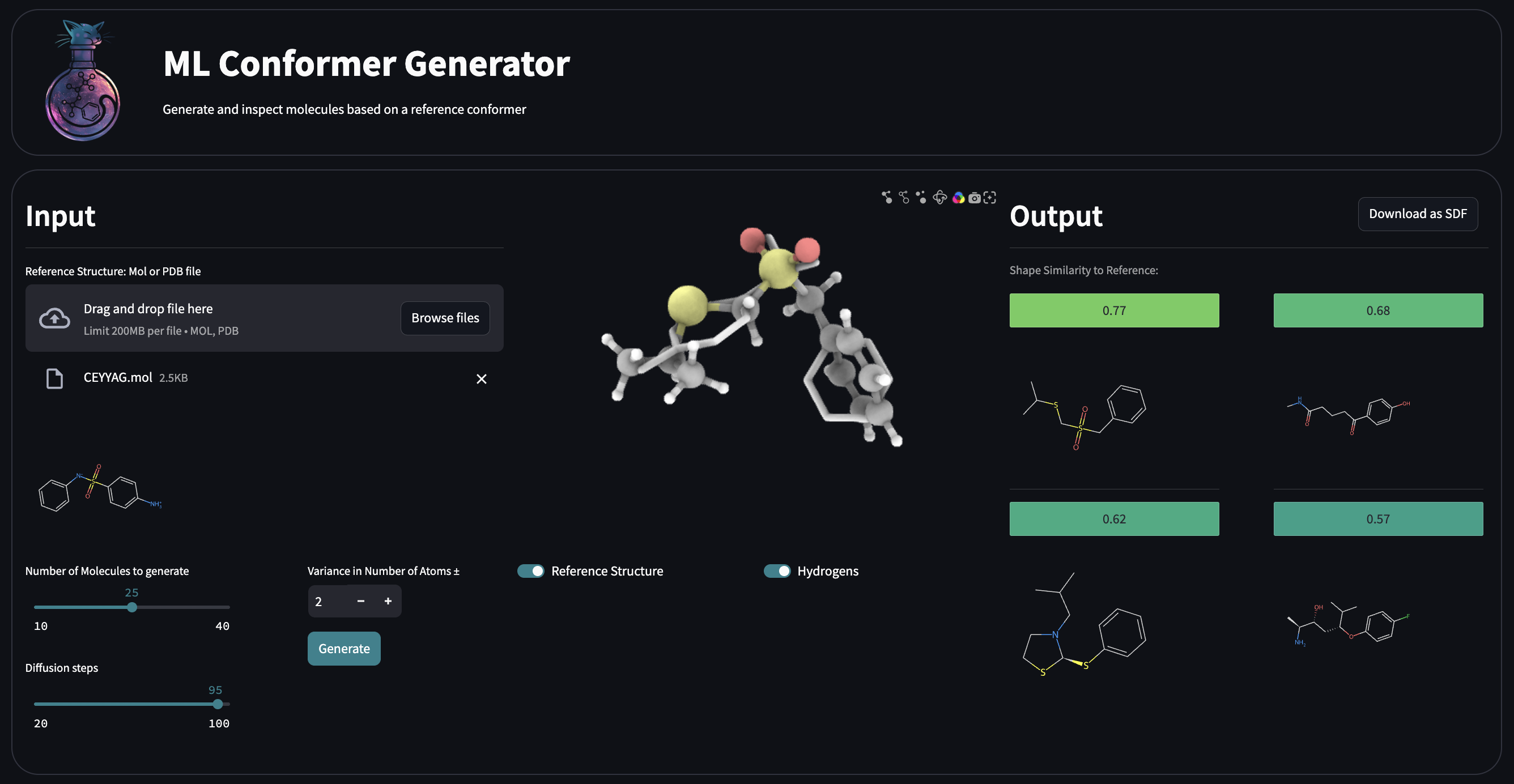
ML Conformer Generator is a tool for spatially-aware molecule generation with an Equivariant Diffusion Model (EDM) and a Graph Convolutional Network (GCN). It is designed to generate 3D molecular conformations that are both chemically valid and spatially similar to a reference shape.
-
Shape-guided molecular generation
Generate novel molecules that conform to arbitrary 3D shapes—such as protein binding pockets or custom-defined spatial regions.
-
Objective-guided Generation
Use reinforcement learning (RL) to steer molecular generation toward higher-scoring candidates, with support for custom scoring functions.
-
Reference-based conformer similarity
Create molecules conformations of which closely resemble a reference structure, supporting scaffold-hopping and ligand-based design workflows.
-
Fragment-based inpainting
Fix specific substructures or fragments within a molecule and complete or grow the rest in a geometrically consistent manner.
-
Inertial Fragment Matching
Generate molecules fragment-by-fragment by leveraging the physical properties of the shape descriptor, improving both shape similarity and chemical validity.
If you use MLConfGen in your research, please cite:
Denis Sapegin, Fedor Bakharev, Dmitry Krupenya, Azamat Gafurov, Konstantin Pildish, and Joseph C. Bear.
Moment of inertia as a simple shape descriptor for diffusion-based shape-constrained molecular generation.
Digital Discovery, 2025.
DOI: 10.1039/D5DD00318K
-
Install the package for your preferred backend:
-
pip install mlconfgen[torch]— use the PyTorch-based inference pipeline -
pip install mlconfgen[onnx]— use the torch-free ONNX runtime version
-
-
Load the weights from Huggingface
edm_moi_chembl_15_39.pt
adj_mat_seer_chembl_15_39.pt
See interactive examples: ./python_api_demo.ipynb
from rdkit import Chem
from mlconfgen import MLConformerGenerator, evaluate_samples
model = MLConformerGenerator(
edm_weights="./edm_moi_chembl_15_39.pt",
adj_mat_seer_weights="./adj_mat_seer_chembl_15_39.pt",
diffusion_steps=100,
)
reference = Chem.MolFromMolFile('./assets/demo_files/ceyyag.mol')
samples = model.generate_conformers(reference_conformer=reference, n_samples=20, variance=2)
aligned_reference, std_samples = evaluate_samples(reference, samples)This solution employs:
- Equivariant Diffusion Model (EDM) [1]: For generating atom coordinates and types under a shape constraint.
- Graph Convolutional Network (GCN) [2]: For predicting atom adjacency matrices.
- Deterministic Standardization Pipeline: For refining and validating generated molecules.
- Trained on 1.6 million compounds from the ChEMBL database.
- Filtered to molecules with 15–39 heavy atoms.
- Supported elements:
H, C, N, O, F, P, S, Cl, Br.
The generated molecules are post-processed through the following steps:
- Largest Fragment picker
- Valence check
- Kekulization
- RDKit sanitization
- Constrained Geometry optimization via MMFF94 Molecular Dynamics
Aligns and Evaluates shape similarity between generated molecules and a reference using Shape Tanimoto Similarity [3] via Gaussian Molecular Volume overlap.
Hydrogens are ignored in both reference and generated samples for this metric.
Tested on 100,000 samples using 1,000 CCDC Virtual Screening [4] reference compounds.
- ⏱ Avg time to generate 50 valid samples: 11.46 sec (NVIDIA H100) (100 samples batch)
- ⚡️ Generation speed: 4.18 valid molecules/sec (100 samples batch)
- 💾 GPU memory (per generation thread): Up to 14.0 GB (
float1639 atoms 100 samples) - 📐 Avg Shape Tanimoto Similarity: 53.32% (Basic generation) - 69.97% (Inertial Fragment Matching)
- 🎯 Max Shape Tanimoto Similarity: 99.69%
- 🔬 Avg Chemical Tanimoto Similarity (2-hop 2048-bit Morgan Fingerprints): 10.87%
- 🧬 % Chemically novel (vs. training set): 99.84%
- ✔️ % Valid molecules (post-standardization): 48% (ML Bond Prediction) - 93% (OpenBabel bond prediction)
- 🔁 % Unique molecules in generated set: 99.94%
- 📎 Fréchet Fingerprint Distance (2-hop 2048-bit Morgan Fingerprints):
- To ChEMBL: 4.13
- To PubChem: 2.64
- To ZINC (250k): 4.95
PoseBusters [5] validity check results:
Overall stats:
- PB-valid molecules: 91.33 %
Detailed Problems:
- position: 0.01 %
- mol_pred_loaded: 0.0 %
- sanitization: 0.01 %
- inchi_convertible: 0.01 %
- all_atoms_connected: 0.0 %
- bond_lengths: 0.24 %
- bond_angles: 0.70 %
- internal_steric_clash: 2.31 %
- aromatic_ring_flatness: 3.34 %
- non-aromatic_ring_non-flatness: 0.27 %
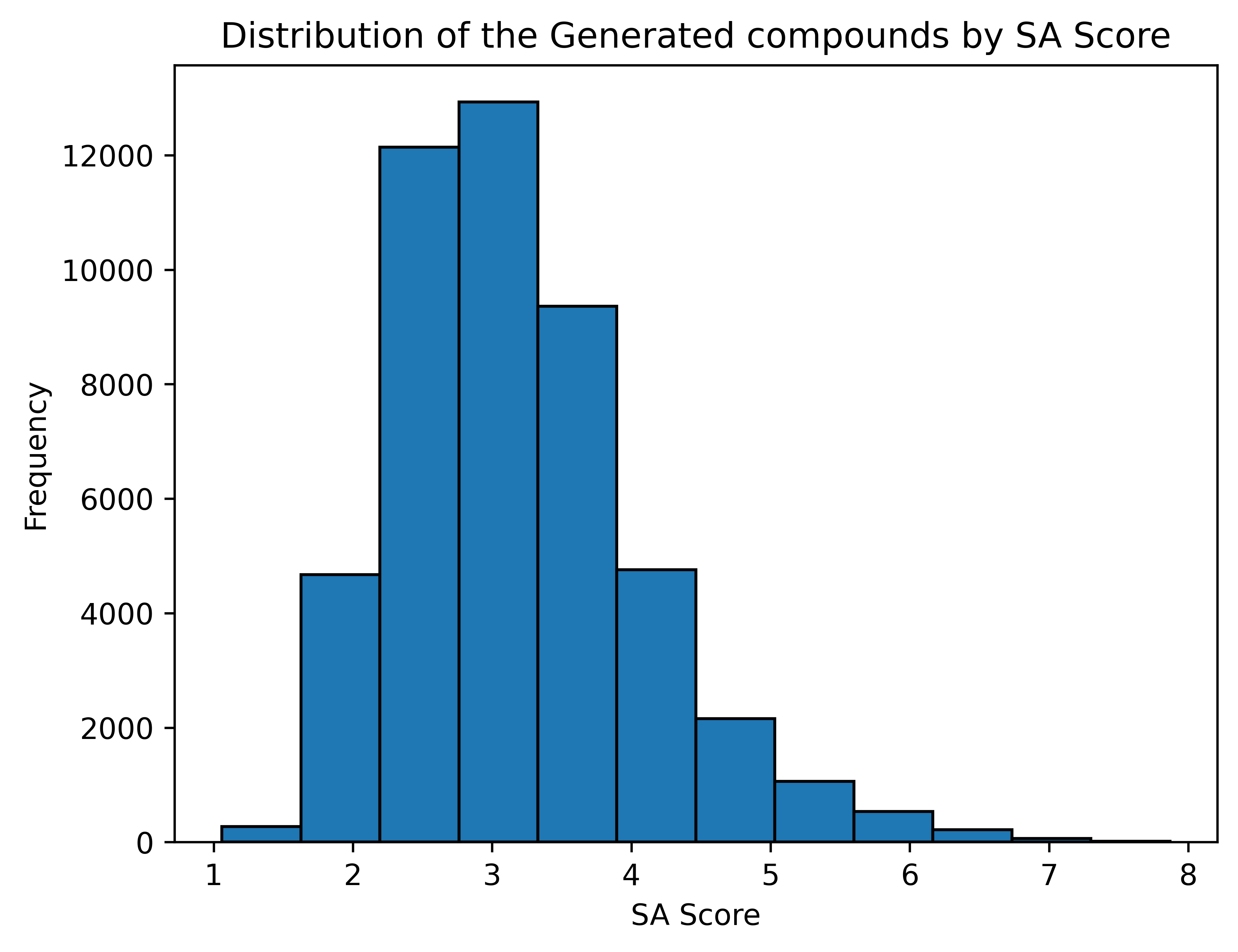
SA Score [6]
1 (easy to make) - 10 (very difficult to make)
Average SA Score: 3.18
MLConformerGenerator supports objective-guided reinforcement learning (RL) fine-tuning, allowing you to steer the generated molecular distribution toward molecules that better match your desired properties.
Scoring functions are fully customizable. The only requirement is that they accept a list of RDKit Mol objects and return a list of scores in the range [0, 1].
A scoring function should follow this interface:
from rdkit import Chem
def scoring_function(mols: list[Chem.Mol | None]) -> list[float]:
...Note
If scoring_function is None, a default scoring function enforcing validity is applied for RL.
from rdkit import Chem
from mlconfgen import MLConformerGenerator
model = MLConformerGenerator(
edm_weights="./edm_moi_chembl_15_39.pt",
adj_mat_seer_weights="./adj_mat_seer_chembl_15_39.pt",
diffusion_steps=10,
)
reference = Chem.MolFromMolFile('./assets/demo_files/ceyyag.mol')
model.fine_tune(
reference_conformer=reference,
variance=1,
n_epochs=20,
sigma=60.0,
lambda_edm_adapter=1.5,
temperature=1.5,
n_samples_per_mol=16,
eval_every=5,
save_dir="./rl_checkpoints"
)
Fine-tuning produces both the best and the latest checkpoints, which can later be loaded into the model:
from mlconfgen import MLConformerGenerator
model = MLConformerGenerator(
edm_weights="./edm_moi_chembl_15_39.pt",
adj_mat_seer_weights="./adj_mat_seer_chembl_15_39.pt",
finetune_checkpoint = "./finetune_checkpoint.pt",
diffusion_steps=10,
)
# Or
model.load_finetune_checkpoint("./finetune_checkpoint.pt")The RL fine-tuning pipeline is compatible with scoring functions from REINVENT4.
If REINVENT4 is installed, you can use ReinventScoreWrapper to load a REINVENT4 scoring configuration and use MLConfGen as a spatially-aware molecule generator.
For working examples, see rl_fine_tuning_demo.ipynb.
from rdkit import Chem
from mlconfgen import MLConformerGenerator
from mlconfgen.rl_fine_tuning.reinvent_score_wrapper import ReinventScoreWrapper
model = MLConformerGenerator(
edm_weights="./edm_moi_chembl_15_39.pt",
adj_mat_seer_weights="./adj_mat_seer_chembl_15_39.pt",
diffusion_steps=10,
)
reference = Chem.MolFromMolFile('./assets/demo_files/ceyyag.mol')
scoring_function = ReinventScoreWrapper("./assets/demo_files/scoring_config.toml")
model.fine_tune(
scoring_function=scoring_function,
reference_conformer=reference,
variance=1,
n_epochs=100,
train_batch_size=128,
eval_batch_size=128,
learning_rate= 8e-5,
sigma=128.0,
lambda_edm_adapter=1.5,
lambda_edm_reg=0.2,
temperature=1.5,
n_samples_per_mol=32,
eval_every=5,
save_dir="./rl_checkpoints_reinvent",
)The Python package and inference code are available on GitHub under Apache 2.0 License
The trained model Weights are available at
And are licensed under CC BY-NC-ND 4.0
The usage of the trained weights for any profit-generating activity is restricted.
For commercial licensing and inference-as-a-service, contact: Denis Sapegin
For torch Free inference an ONNX version of the model is present.
Weights of the model in ONNX format are available at:
egnn_chembl_15_39.onnx
adj_mat_seer_chembl_15_39.onnx
from mlconfgen import MLConformerGeneratorONNX
from rdkit import Chem
model = MLConformerGeneratorONNX(
egnn_onnx="./egnn_chembl_15_39.onnx",
adj_mat_seer_onnx="./adj_mat_seer_chembl_15_39.onnx",
diffusion_steps=100,
)
reference = Chem.MolFromMolFile('./assets/demo_files/yibfeu.mol')
samples = model.generate_conformers(reference_conformer=reference, n_samples=20, variance=2)Install ONNX GPU runtime (if needed):
pip install onnxruntime-gpu
An option to compile the model to ONNX is provided
requires onnxscript==0.2.2
pip install onnxscript
from mlconfgen import MLConformerGenerator
from onnx_export import export_to_onnx
model = MLConformerGenerator()
export_to_onnx(model)This compiles and saves the ONNX files to: ./
To execute all tests (including slow generation ones)
pytest -v tests
To bypass generation tests
pytest -v tests -m "not slow"
- Move the trained PyTorch weights into
./streamlit_app
./streamlit_app/edm_moi_chembl_15_39.pt
./streamlit_app/adj_mat_seer_chembl_15_39.pt
-
Install the dependencies
pip install -r ./streamlit_app/requirements.txt -
Bring the app UI up:
cd ./streamlit_app streamlit run app.py
-
To enable development mode for the 3D viewer (
stspeck), set_RELEASE = Falsein./streamlit/stspeck/__init__.py. -
Navigate to the 3D viewer frontend and start the development server:
cd ./frontend/speck/frontend npm run startThis will launch the dev server at
http://localhost:3001 -
In a separate terminal, run the Streamlit app from the root frontend directory:
cd ./streamlit_app streamlit run app.py -
To build the production version of the 3D viewer, run:
cd ./streamlit_app/stspeck/frontend npm run build